COVID-19 Sounds App
< Research Updates
COVID-19 progression prediction: A longitudinal study
June 21, 2022 — Can we continuously detect and track individuals’ COVID-19 disease progression in a remote monitoring context? While previous studies have shown that voice and respiratory sounds are promising for COVID-19 detection, these approaches only focus on one-off detection given snapshots of available data, but do not continuously monitor COVID-19 disease progression, especially recovery, through longitudinal audio data. Tracking disease progression characteristics and patterns of recovery could lead to more timely treatment or treatment adjustment, as well as better resource management in healthcare systems.
The primary objective of this study is to explore the potential of longitudinal audio samples over time for COVID-19 disease progression prediction and, especially, recovery trend prediction using sequential deep learning. The changes in longitudinal audio samples, referred to as audio dynamics, are associated with COVID-19 disease progression, thus modelling the audio dynamics can potentially capture the underlying disease progression process and further aid COVID-19 progression prediction. The results are described in a research article published in JMIR, the leading journal for digital medicine and health.
Method
We selected a subset of 212 individuals for analysis, who provided respiratory audio data (i.e., breathing, cough, and voice) and corresponding test results for a period of 5 to 385 days. We developed and validated a deep learning-enabled method using Gated Recurrent Units (GRUs) to detect COVID-19 disease progression, by exploring the audio dynamics of individuals’ historical audio biomarkers.
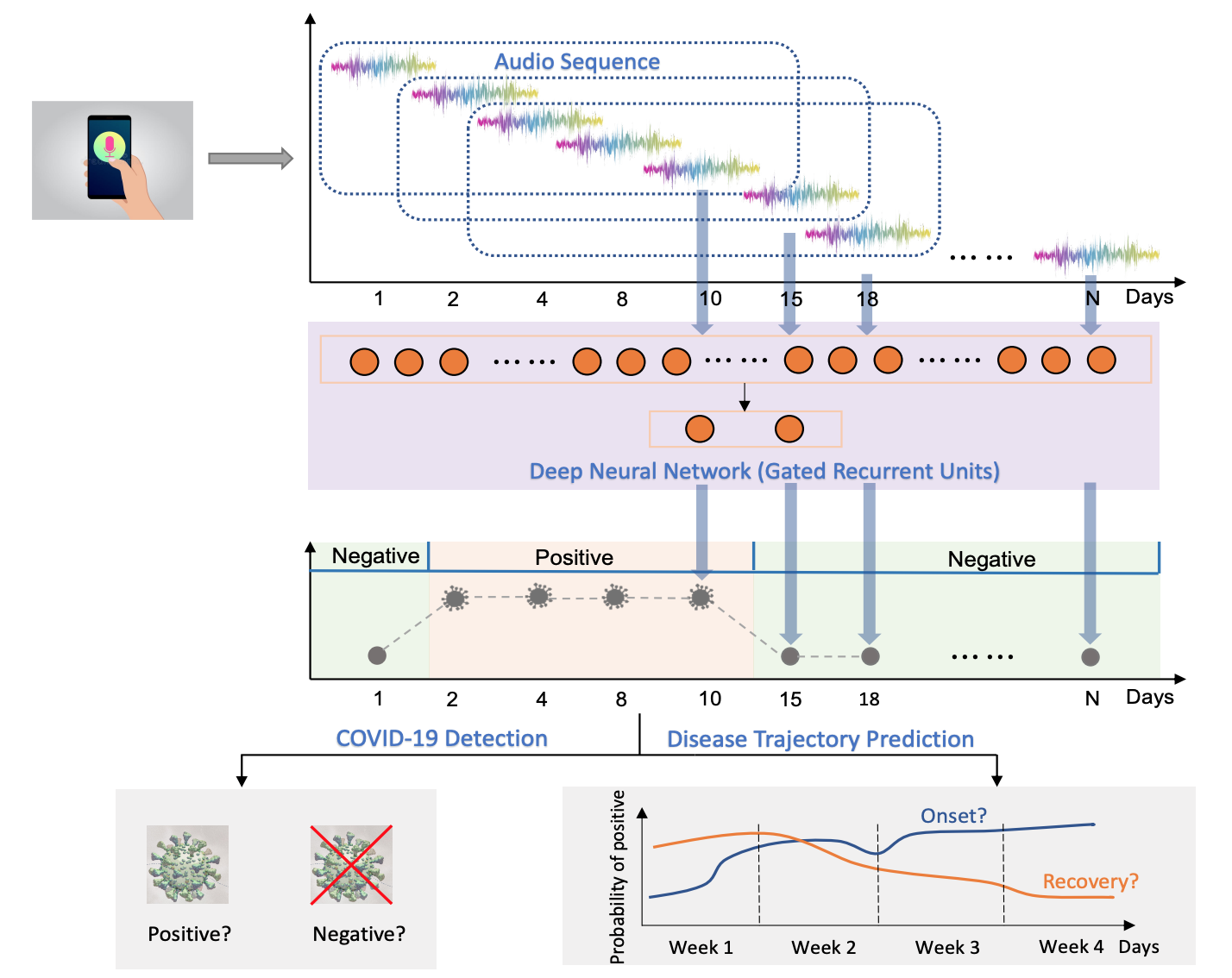
We investigated two tasks: i) COVID-19 detection that validates whether capturing audio dynamics over time can aid in distinguishing between COVID-19 positive and negative (healthy) audio samples; ii) disease trajectory prediction that verifies whether the model can successfully track an individuals’ disease progression by comparing the predicted trajectory with the self-reported labels.
The proposed model consists of a pre-trained convolutional network named VGGish [1] to extract the high-level audio information, and GRUs to capture the temporal dependencies of the longitudinal audio samples for COVID-19 progression detection. As the voice data contains 8 different languages, the different disease prevalence for each language could lead to the model recognising the language instead of COVID-19 related information, e.g. classifying Italian speakers as positive and English as negative owing to higher prevalence in Italian-speaking users and lower prevalence in English-speakers. Therefore, a multi-task learning framework was proposed with language recognition as the auxiliary task to reduce the language bias.
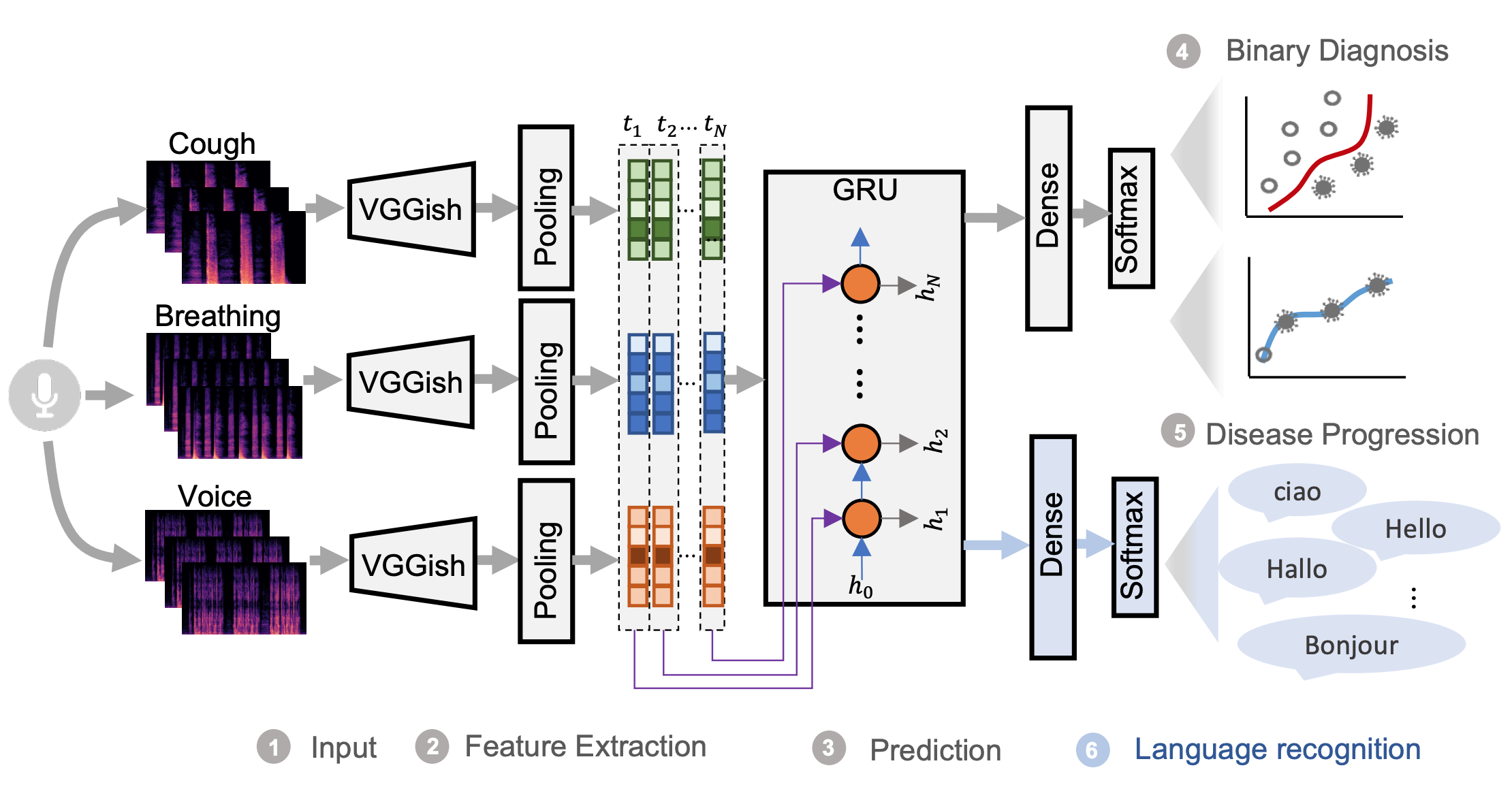
Findings
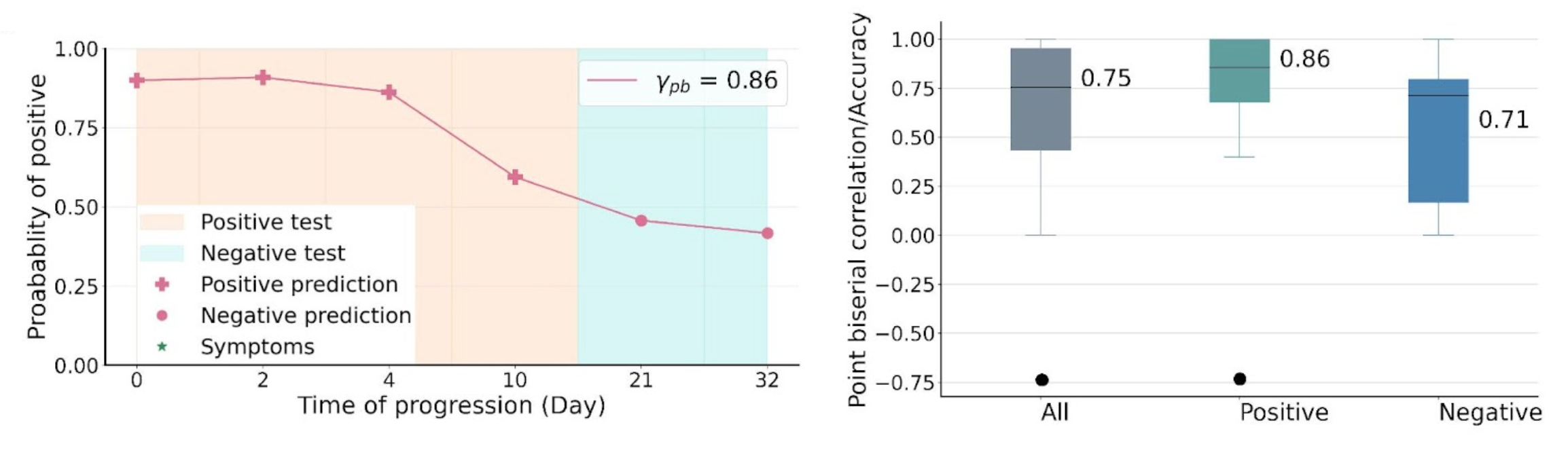
The proposed system yielded a strong performance in distinguishing between COVID-19 positive and negative audio samples with AUC-ROC of 0.79, sensitivity of 0.75 and specificity of 0.71, significantly outperforming SOTA approaches that do not leverage temporal dynamics. We further examined the predicted disease progression trajectory, with an example of one participant’s disease progression and prediction shown below. The predictions display high consistency with the longitudinal test results, with high correlation of 0.86. Overall, the system yielded a correlation of 0.75 in the test cohort, and 0.86 and 0.71 for positive and negative groups respectively. For a subset of the test cohort who report disease recovery, the system yielded 0.86, suggesting its potential in home monitoring during postpeak and postpandemic time.
Contribution
To the best of our knowledge, this is the first work exploring the longitudinal audio dynamics for COVID-19 disease progression prediction. This framework provides a flexible, affordable and timely tool for COVID-19 disease tracking, and more importantly, it also provides a proof of concept of how telemonitoring could be applicable to respiratory diseases monitoring, in general. Our future work includes audio-based COVID-19 disease progression forecasting ahead of time, and model development for personalised disease progression tracking. The open-source code for this paper will be released on Github soon.
How to cite our paper
Ting Dang, Jing Han*, Tong Xia*, Dimitris Spathis, Erika Bondareva^, Chloë Siegele-Brown^, Jagmohan Chauhan^, Andreas Grammenos^, Apinan Hasthanasombat^, Andres Floto, Pietro Cicuta and Cecilia Mascolo. "Exploring Longitudinal Cough, Breath, and Voice Data for COVID-19 Disease Progression Prediction via Sequential Deep Learning: Model Development and Validation". Journal of medical Internet research (2022)
[PDF].
*equal contribution, ^alphabetical order
[1] Shawn Hershey, Sourish Chaudhuri, et al. CNN architectures for large-scale audio classification. In Proceedings of the International Conference on Acoustics, Speech, and Signal Processing (ICASSP), pages 131–135, 2017
